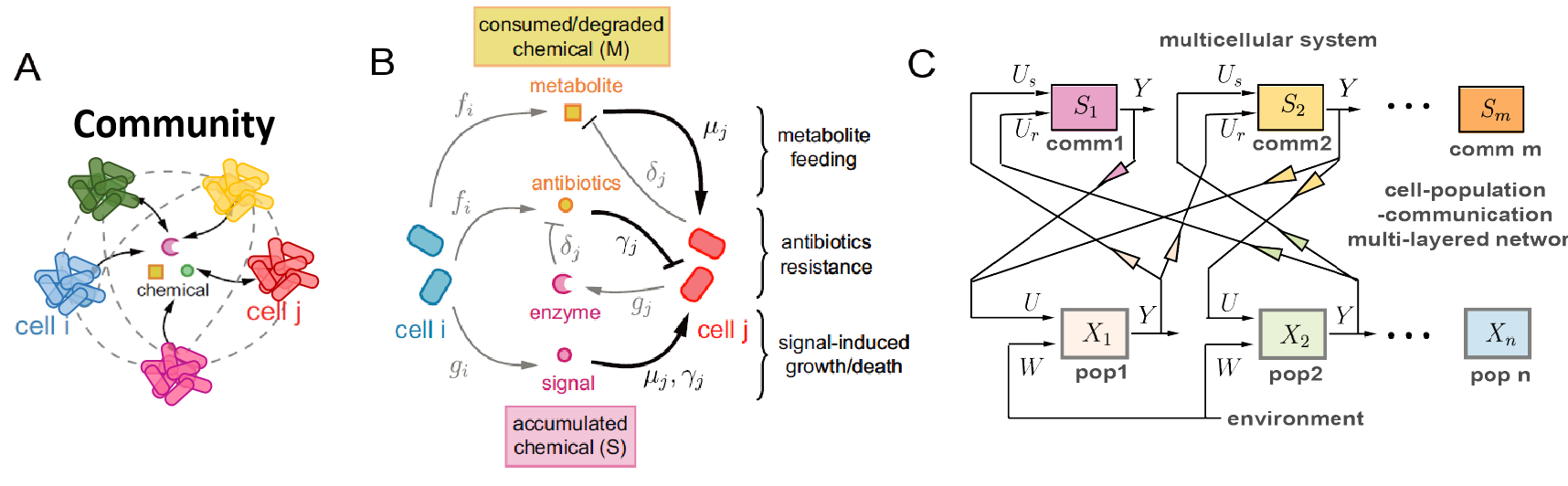
The spatial-temporal evolution of interspecies interactions in multicellular communities plays an important role for them to survive, adapt and perform functions in certain environments. On the one hand, interspecies interactions facilitate self-organized spatial structures. For example, cell populations grow or actively migrate towards nutrients through cross-feeding interactions. Interspecies killing also restructures a community via contact-based inhibition or diffusible toxins. On the other hand, the efficiency of interspecies interactions is strongly impacted by the physical location and distance of cells in defined spatial environments. Thererfore, the interspecies interactions and spatial environments co-evolve to maintain long-term stability. We are particularly interested in why certain spatial structures in multicellular communities provide benefits for cells to survive changing environments and how cell populations self-organize into such spatial structures. By investigating the mechanisms of the spatial-temportal evolution of interspecies interactions, we are able to engineer synthetic circuits that perform diverse interactions and achieve stable coexistence and robustness to environmental disturbances. tail Chemical-mediated interactions among microbial populations. A. Sketech of a chemical-mediated cell-cell interaction network. B. Intereactions with detailed mechanisms depending on if the signaling chemcal is consumed/degraded by receiver cells or accumulates. The chemical synthesis rate from cell type i is denoted by index i, and the chemical consumption rate by cell type j is denoted by index j. C. A bipartite structure of microbial interaction network that includes a population module and a communication module. The population module senses signaling chemicals as input (U) and the output is population-dependent chenical synthesis or comsumption (Y). The communication module takes in inputs from sender (Us) and receiver (Ur) cell populations.
For example, interspecies interactions can be mediated via chemicals, including small signaling molecules, metabolites, toxins, etc. Based on detailed mechanisms of signal synthesis, secretion, diffusion, uptake and their effects on receiver cells, we are able to build up signaling modules with input from sender cells and output from receiver cells. These signaling modules may exhibit diver input-output responses and can characterize how cell populations communicate with each other. Therefore, we porpose a bipartite structure of population-communication networks that can be easily upscaled to multiple cell populations and communication signaling channels. Compared to pairwise interaction network structure, now we have a more mechanistic description of interactions and a more quantitative characterization of population dynamics.